Countdown to 1 million
The recent August update to the Cambridge Structural Database (CSD) brought the total number of entries in the database to over 950,000, meaning the next big milestone will be 1 million. This is a huge achievement of the crystallographic community, and in the months leading up to this milestone we’ll be demonstrating the value that can be gained from this crystal data and looking to what can be accomplished in the future.
Update 24/01/2025: Since this blog was written, the Cambridge Structural Database (CSD) has surpassed 1 million. For more up-to-date information;
- Learn more about the CSD here.
- Register for our monthly email newsletter here to get the latest news.
Now that we’re really starting to get close to this nice round million figure, it’s probably worth considering more carefully what we’re counting to get to 1 million. This is a question that is raised quite regularly, and the short answer to “What is the size of the CSD?” is given in one of our FAQs. We’ve been thinking in quite general terms about the ‘CSD 1 million’ for a while now, and I’ve written blogs in the past commenting on the growth of the CSD when we issued CCDC 900,000 and released the 700,000th CSD entry, but there are a few things to consider in our count.
Firstly, and hopefully this won’t come as a surprise, we probably already have passed the million mark. We, and most major journal publishers such as the RSC, recommend crystallographic data is deposited at the CCDC as part of the article submission process. This data is then available for journal referees, but it is otherwise kept confidentially until the researchers publish their work, either as an article or directly via a CSD Communication. Therefore, the first criterion for our count is one million structures freely available to the scientific community through the CSD.
The next question to consider is what we want to count. The CSD has grown and evolved since its inception in 1965, and currently an individual CSD entry refers to one published report of a crystal structure. At the time of writing, there are just over 970,000 CSD entries, and the picture below shows one of these entries; CSD refcode HEFVEW01, from the latest August CSD update.

CSD refcode HEFVEW01, also known as CSD refcode HEFVEW, which corresponds to the single crystal structure https://dx.doi.org/10.5517/ccdc.csd.cc1nx5zq
If we also look for the entry with CSD refcode HEFVEW you’ll see this is the same crystallographic data that has been reported by the authors in two separate publications, so we have two CSD entries and one unique crystal structure. This isn’t a particularly uncommon occurrence, as research projects develop they continue to build on previous findings and crystallographic data may continue to be relevant in subsequent publications.
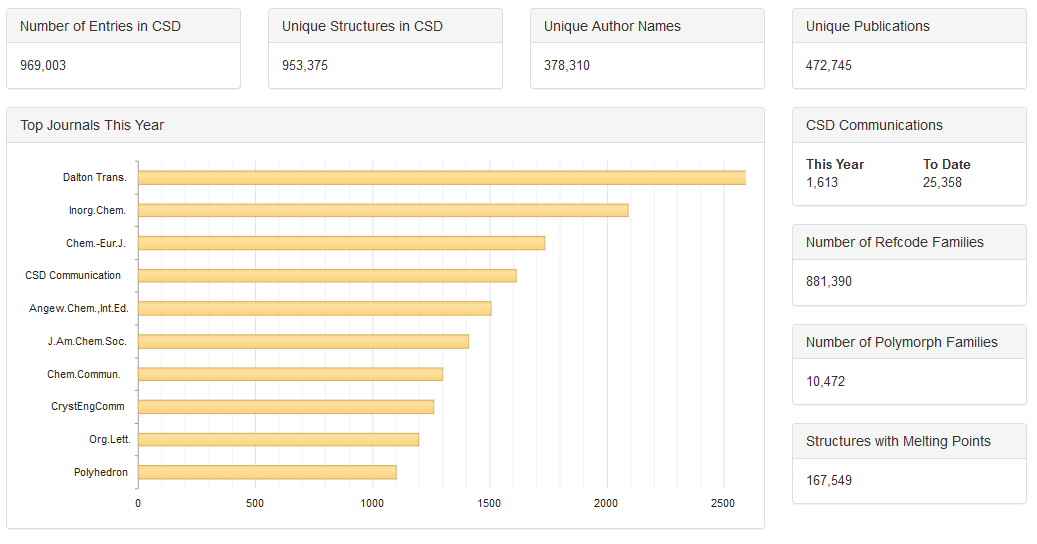
Part of our new CSD Statistics page (/CCDCStats/) , showing the number of CSD entries and datasets.
If you’re curious just how often this occurs, we have recently released a new CSD Statistics page, which shows just that! The CSD currently holds 970,693 entries, and these come from 955,017 unique crystal structures – moving us slightly further away from our 1 million. Another point to note here is that broadly speaking, determining a crystal structure has two steps; the diffraction data must be collected, and secondly a structural model is created and refined to best fit the collected data. Occasionally the same data may be interpreted in different ways, creating a different structural model – the eminent crystallographer Richard Marsh was famous for this sort of re-analysis. Therefore, when we refer to a ‘unique crystal structure’, we really mean a unique combination of data collection and refinement model. Our statistics page also considers one other issue – any data from a publication that has subsequently been retracted is not included in our totals.

CSD refcodes PROLIN03 and PROLIN04, two polymorphs of the compound L-Proline
A final issue to consider is we could perhaps come up with an even more strict definition of 1 million – we could wait until we have 1 million unique chemical structures. If we look at the amino acid L-Proline we can see there is a CSD refcode family PROLIN-PROLIN05, containing six unique crystal structures. These range from the first structure reported (CSD refcode PROLIN) in 1965, a powder diffraction structure in 2010 (CSD refcode PROLIN01) to the discovery of a second polymorph from a synchrotron powder diffraction experiment earlier this year (CSD refcode PROLIN04). All six entries contain useful information, but they are all of the same chemical composition. It’s easy for us to do this calculation too, because we organise the CSD with refcode families which contain all instances of the same chemical composition. This would give us a significantly smaller number; as you can see on our CSD Statistics page there are 882,855 refcode families in the CSD. We feel this definition is a bit too strict however, reports of different polymorphs of a structure, or the same polymorph at different temperatures and pressures for example, all provide valuable data and insights to the scientific community. So next year when we’ll be celebrating ‘CSD 1 million’, hopefully it will be clear that the achievement we’re celebrating is that worldwide crystallographers will have produced 1 million unique crystal structures that are available for the community, something we should all be very proud of.
As we begin our countdown to CSD 1 million, do let us know if you have any thoughts or feedback! As always, we are available via our website or through our CCDC social media channels (Facebook, LinkedIn and X or Twitter). Do check our channels (and search for #CSD1Million) in the coming months as we prepare for the celebrations!
Tags