CSDU: On-Demand Learning
CSDU is a series of on-demand training modules where U can learn how to harness cheminformatics and data insights from the CSD to use in your work.
Here you can learn about:
- The Cambridge Structural Database (CSD).
- How to use the CSD software and data to inform and advance your research.
- How cheminformatics and structural chemistry can be applied to problems in the discovery and development of new molecules and materials.
Learn more about how our CSDU modules work in this blog.
Upon completion of these modules, you will be awarded a certificate or LinkedIn badge certification to showcase your achievement.
Sign up for our Training, Education and Outreach newsletter
Who can use CSDU?
Everybody! From beginners to advanced users, we hope to grow the CSDU collection to cover a wide range of applications.
Note: you will need a licence to complete the “try it yourself” exercises. Ask your site administrator, or contact our team here if you need to request a licence.
How does CSDU work?
Choose a module below, and work through the steps shown. Modules are fully on-demand, so you can work at your own pace.
All modules follow the basic format:
- U Watch – get context and see examples from our tutors in an online video.
- U Try – work through the example yourself, following the tutorial worksheet.
- U Test – check your knowledge in an online quiz, and get your certificate of completion or LinkedIn badge certification by email.
CSDU Modules
Start your CSDU courses with the modules below!
We regularly add new CSDU modules. Register for our newsletter so you don’t miss new releases!
Entry Level
CSDU modules in this section do not require any previous knowledge of the CSD software. Start from here!
Searching the CSD 101 - Searching Structures Online with WebCSD
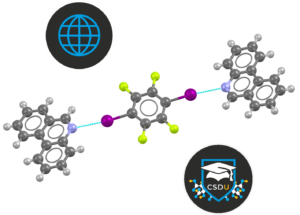
WebCSD allows you to search for structures in the Cambridge Structural Database (CSD) and to find the context of the structure in relation to its originality and/or comparison to other known molecules. WebCSD enables you to run different query types and it returns the most up-to-date possible results. Original deposited data can be obtained and exported for further analysis as well as the values of 3D parameters defined in a structure search.
In this module, you will explore advanced functionality in WebCSD, with focus on the Structure Search features.
Pharmacophore Searching 101 - Introduction to CSD-CrossMiner
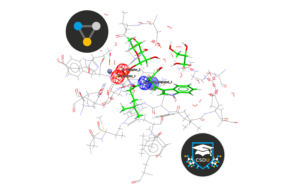
CSD-CrossMiner is software that allows you to build pharmacophore queries and simultaneously mine the Cambridge Structural Database (CSD), the Protein Data Bank (PDB), and your proprietary structural data to quickly identify off-target effects, alternative scaffolds, similar binding sites, interaction motifs, bioisosteres, and more.
In this CSDU module, we will focus on the basics of performing a pharmacophore search using CSD-CrossMiner. Those that complete this module and answer the final quiz questions correctly will gain a LinkedIn badge certification to showcase their achievement.
Intermediate Level
CSDU modules for next level structural analysis.
Advanced Level
CSDU modules requiring fluency with CSD Software.
More modules under development
Reply to our newsletter to let us know what modules you would like to see next…
Sign up for our Training, Education and Outreach newsletter
Receive updates about training events and resources.





